How is mRNA different than DNA?
Firstly the structure is quite different, DNA is huge and has Two strands (also double helix), while mRNA is small single stranded. mRNA also carries the code for DNA, DNA has all the genetic codes in its bases, mRNA gets the code from DNA during transcription and eventually carriers the code to produce amino acids and proteins. A bunch of other differences is DNA uses the sugar Deoxyribose and mRNA uses sugar ribose. DNA is formed in DNA replication, mRNA is from transcription. DNA is found in the nucleus while mRNA is mostly found in the cytoplasm.
Process of Transcription:
Unwinding and Unzipping
So the DNA Helicase breaks the H-bonds in the DNA allowing mRNA to complimentary base pair with the DNA.
Complimentary base pairing
The RNA polymerase H-bonds together with floating mRNA with the DNA. The mRNA builds up and produces a polypeptide of mRNA. The DNA Information is copied onto the mRNA.
Separation
The mRNA separates from the DNA. Since the DNA is too big and important to leave the Nucleus, the mRNA a smaller single stranded polypeptide leaves with the DNA’s Information in the Nucleus pores and enters the cytoplasm so it can be read by the ribosome to create proteins.
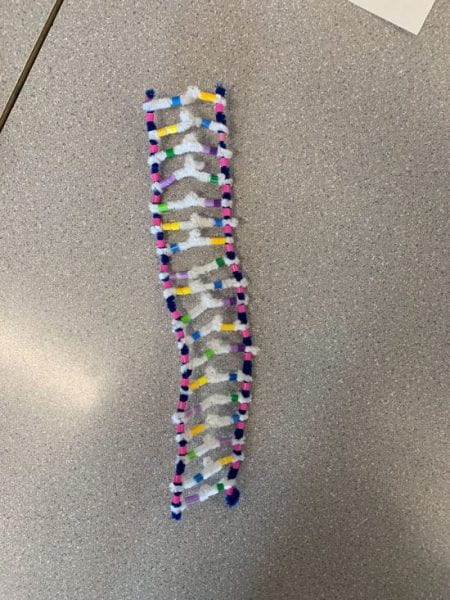
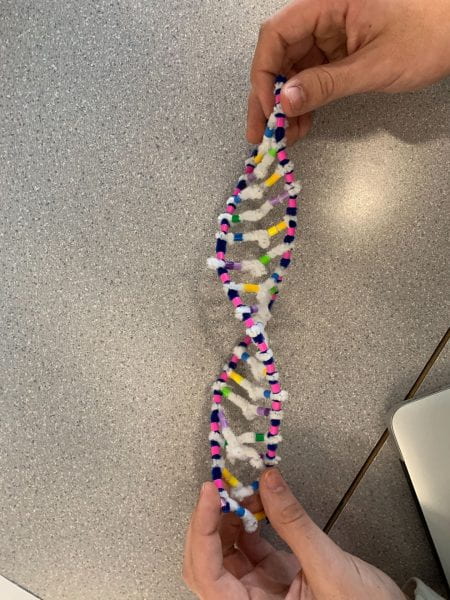
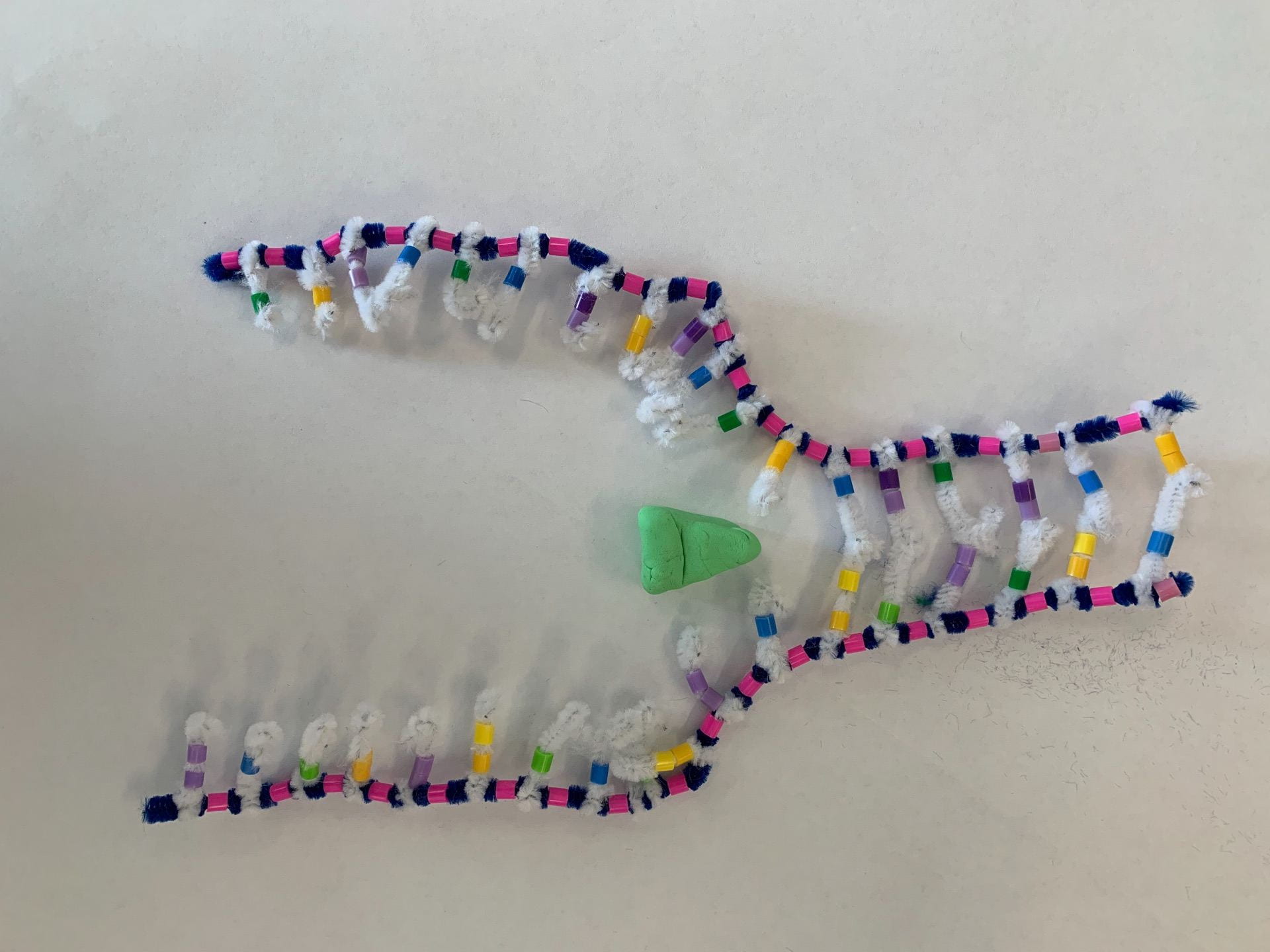
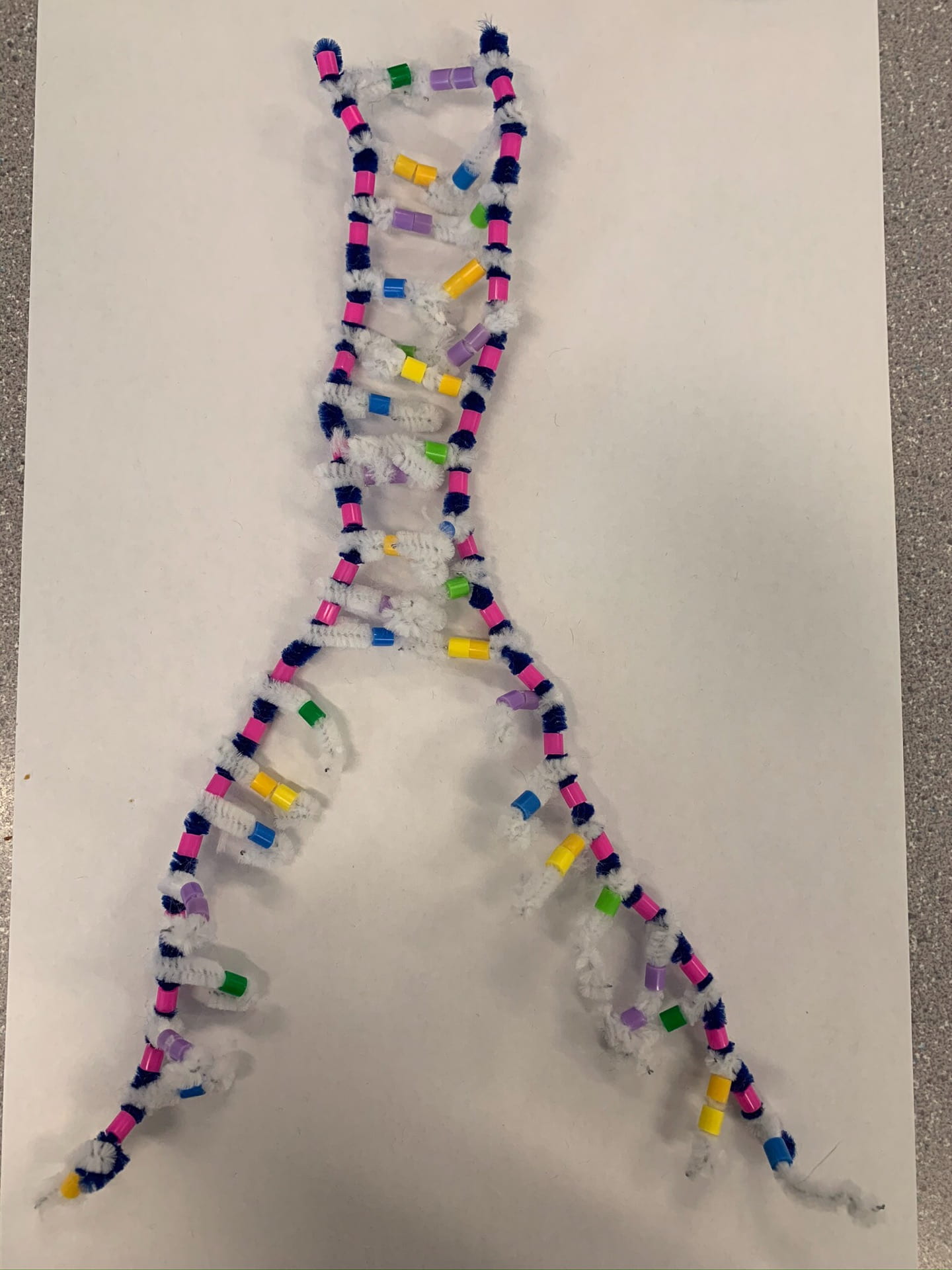
How did this model do a good job in modelling RNA transcription? What ways is it inaccurate?
like the pipe cleaner activity it shows the overall process, but it’s hard to show every small detail and process. An example is we cannot see RNA Polymerase really do its job as it attaches single nucleotides and then bonds them next to each other for the backbone, but in this activity instead it shows it attaching in one step.
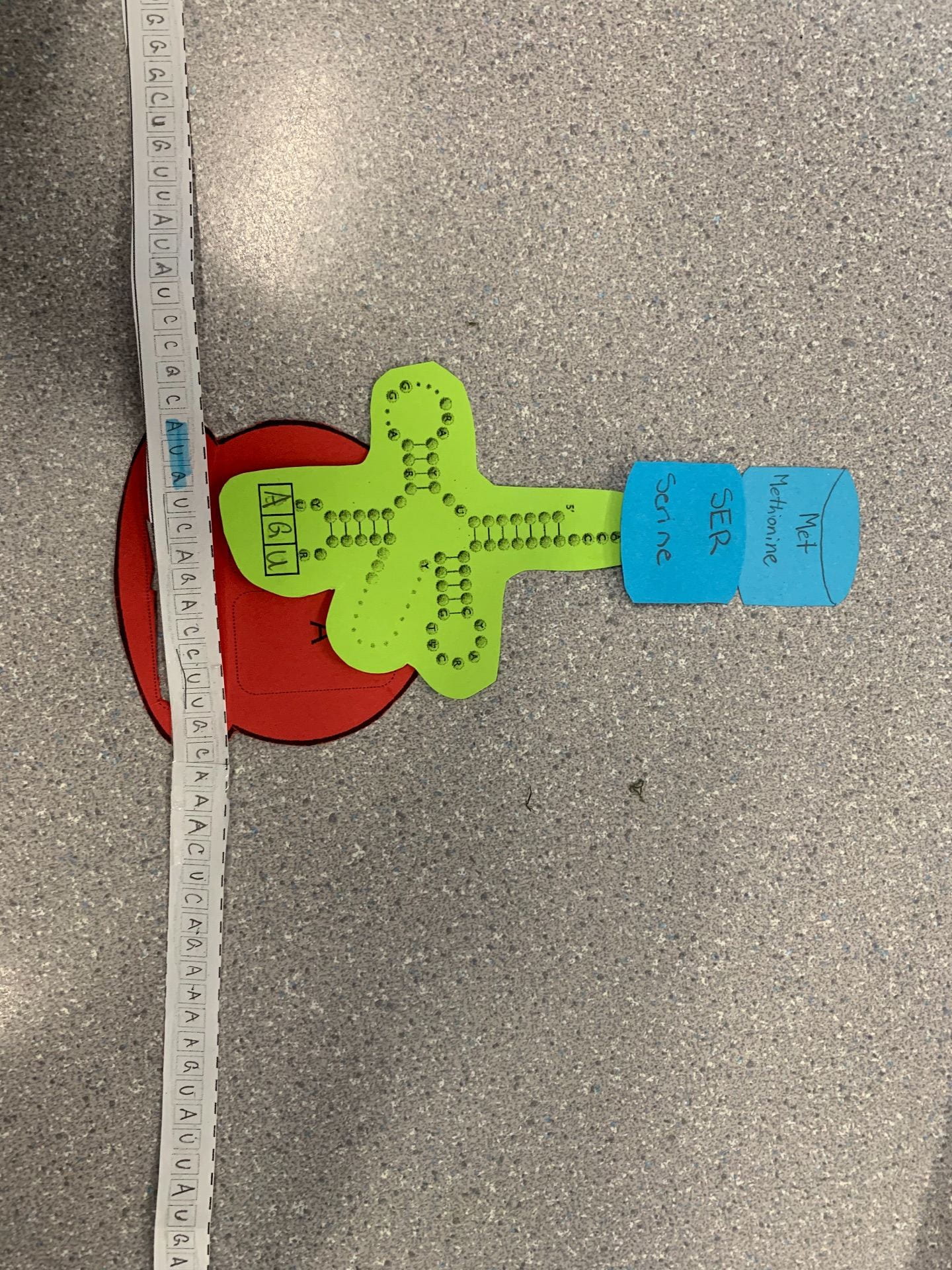
Describe the process of translation: initiation, elongation, and termination:
Initiation
this starts the translation process, the P sight of the Ribosome reads the AUG start Codon on the mRNA strand. The tRNA will bring in the first amino acid then it will start elongation until a stop codon is reached. 
Elongation
The amino acid chain starts to grow. A-site reads the next codon on the mRNA strand and then brings the matching tRNA. Then the tRNA on the P-site transfers over to the tRNA on the A-site.
Termination
A stop Codon is read on the mRNA strand, and this tells the ribosome to stop reading, and it produces no amino acid. Then the Ribosome releases the amino acid chain.
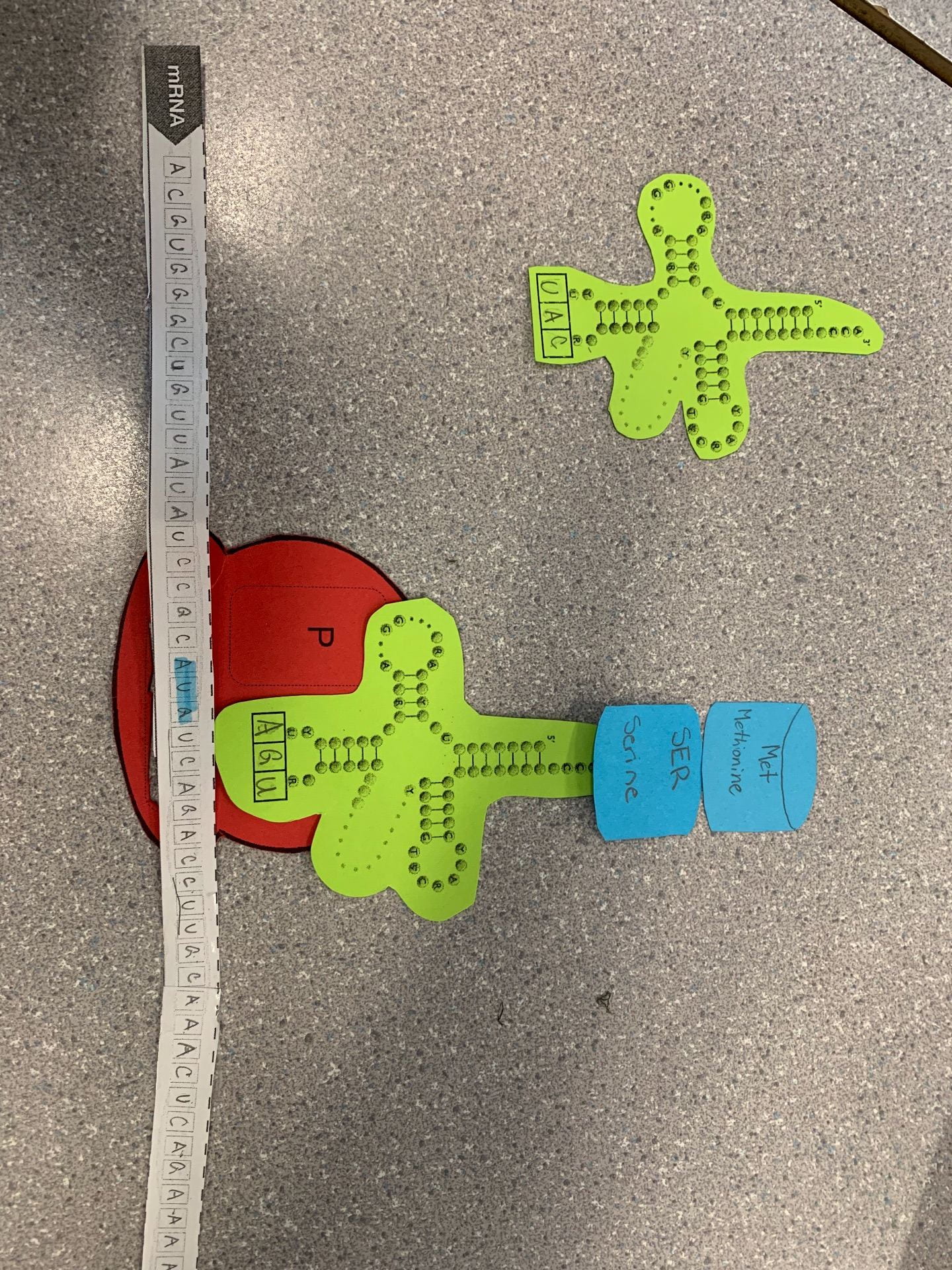
How does today’s activity do a good job of modelling the process of translation? In what ways was our model inaccurate?
It’s accurate showing the Ribosome holding on to the RNA. Also the tRNA building process. amino acid in the p-site and a-site match up with the codons. However what is inaccurate is the amino acid in the P-site and A-site match up with the codons.